Processing
scmdata has some support for processing ScmRun instances to calculate statistics
of interest. Here we provide examples of how to use them.
At present, we can calculate:
crossing times (e.g. 1.5C crossing times)
crossing time quantiles
exceedance probabilities
peak
peak year
categorisation in line with SR1.5
a set of summary variables
Load some data
For this demonstration, we are going to use MAGICC output from [RCMIP Phase 2] as available at https://zenodo.org/record/4624566/files/data-processed-submission-database-hadcrut5-target-MAGICCv7.5.1.tar.gz?download=1. Here we have just extracted the air temperature output for the SSPs from 1995 to 2100.
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import seaborn as sns
import scmdata.processing
from scmdata import ScmRun
/tmp/ipykernel_836/579853008.py:3: DeprecationWarning:
Pyarrow will become a required dependency of pandas in the next major release of pandas (pandas 3.0),
(to allow more performant data types, such as the Arrow string type, and better interoperability with other libraries)
but was not found to be installed on your system.
If this would cause problems for you,
please provide us feedback at https://github.com/pandas-dev/pandas/issues/54466
import pandas as pd
/home/docs/checkouts/readthedocs.org/user_builds/scmdata/checkouts/stable/src/scmdata/database/_database.py:9: TqdmWarning: IProgress not found. Please update jupyter and ipywidgets. See https://ipywidgets.readthedocs.io/en/stable/user_install.html
import tqdm.autonotebook as tqdman
magicc_output = ScmRun("magicc-rcmip-phase-2-gsat-output.csv")
magicc_output
<ScmRun (timeseries: 4800, timepoints: 106)>
Time:
Start: 1995-01-01T00:00:00
End: 2100-01-01T00:00:00
Meta:
climate_model ensemble_member model region scenario unit \
0 MAGICCv7.5.1 0 unspecified World ssp585 K
1 MAGICCv7.5.1 1 unspecified World ssp585 K
2 MAGICCv7.5.1 2 unspecified World ssp585 K
3 MAGICCv7.5.1 3 unspecified World ssp585 K
4 MAGICCv7.5.1 4 unspecified World ssp585 K
... ... ... ... ... ... ...
4795 MAGICCv7.5.1 595 unspecified World ssp119 K
4796 MAGICCv7.5.1 596 unspecified World ssp119 K
4797 MAGICCv7.5.1 597 unspecified World ssp119 K
4798 MAGICCv7.5.1 598 unspecified World ssp119 K
4799 MAGICCv7.5.1 599 unspecified World ssp119 K
variable
0 Surface Air Temperature Change
1 Surface Air Temperature Change
2 Surface Air Temperature Change
3 Surface Air Temperature Change
4 Surface Air Temperature Change
... ...
4795 Surface Air Temperature Change
4796 Surface Air Temperature Change
4797 Surface Air Temperature Change
4798 Surface Air Temperature Change
4799 Surface Air Temperature Change
[4800 rows x 7 columns]
Crossing times
The first thing we do is show how to calculate the crossing times of a given threshold.
crossing_time_15 = scmdata.processing.calculate_crossing_times(
magicc_output,
threshold=1.5,
)
crossing_time_15
climate_model ensemble_member model region scenario unit variable
MAGICCv7.5.1 0 unspecified World ssp585 K Surface Air Temperature Change 2025.0
1 unspecified World ssp585 K Surface Air Temperature Change 2029.0
2 unspecified World ssp585 K Surface Air Temperature Change 2024.0
3 unspecified World ssp585 K Surface Air Temperature Change 2023.0
4 unspecified World ssp585 K Surface Air Temperature Change 2023.0
...
595 unspecified World ssp119 K Surface Air Temperature Change 2023.0
596 unspecified World ssp119 K Surface Air Temperature Change NaN
597 unspecified World ssp119 K Surface Air Temperature Change 2026.0
598 unspecified World ssp119 K Surface Air Temperature Change 2019.0
599 unspecified World ssp119 K Surface Air Temperature Change 2024.0
Length: 4800, dtype: float64
The output is a pd.Series, which is useful for many other pieces of work.
For example, we could make a plot with e.g. seaborn.
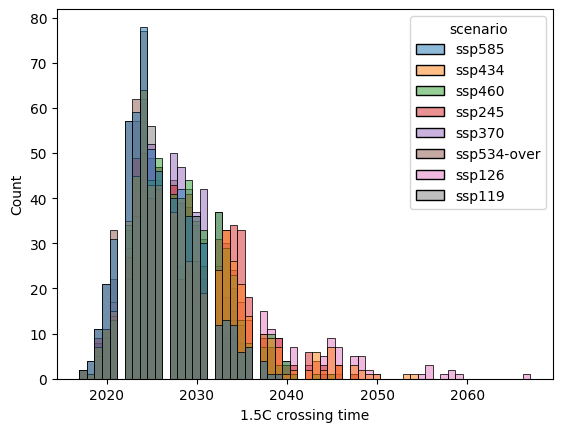
label = "1.5C crossing time"
pdf = crossing_time_15.reset_index().rename({0: label}, axis="columns")
sns.histplot(data=pdf, x=label, hue="scenario")
<Axes: xlabel='1.5C crossing time', ylabel='Count'>
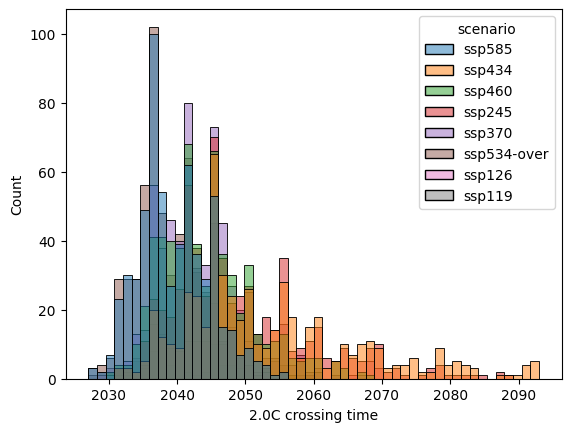
label = "2.0C crossing time"
crossing_time_20 = scmdata.processing.calculate_crossing_times(
magicc_output,
threshold=2.0,
)
pdf = crossing_time_20.reset_index().rename({0: label}, axis="columns")
sns.histplot(data=pdf, x=label, hue="scenario")
<Axes: xlabel='2.0C crossing time', ylabel='Count'>
Crossing time quantiles
Calculating the quantiles of crossing times is a bit fiddly because some ensemble members will not cross the threshold at all. We show how quantiles can be calculated sensibly below. The calculation will return nan if that quantile in the crossing times corresponds to an ensemble member which never crosses the threshold.
scmdata.processing.calculate_crossing_times_quantiles(
crossing_time_15,
groupby=["climate_model", "model", "scenario"],
quantiles=(0.05, 0.17, 0.5, 0.83, 0.95),
)
climate_model model scenario quantile
MAGICCv7.5.1 unspecified ssp119 0.05 2021.00
0.17 2023.00
0.50 2027.00
0.83 NaN
0.95 NaN
ssp126 0.05 2021.00
0.17 2023.00
0.50 2028.00
0.83 2038.17
0.95 NaN
ssp245 0.05 2021.00
0.17 2023.00
0.50 2028.00
0.83 2034.00
0.95 2039.00
ssp370 0.05 2021.00
0.17 2024.00
0.50 2028.00
0.83 2032.00
0.95 2036.00
ssp434 0.05 2021.00
0.17 2023.00
0.50 2027.00
0.83 2034.00
0.95 2041.00
ssp460 0.05 2021.00
0.17 2023.00
0.50 2027.00
0.83 2032.00
0.95 2036.00
ssp534-over 0.05 2020.00
0.17 2022.00
0.50 2025.00
0.83 2030.00
0.95 2033.05
ssp585 0.05 2020.00
0.17 2022.00
0.50 2025.00
0.83 2030.00
0.95 2034.00
dtype: float64
In the above, we can see that the 83rd percentile of crossing times is in fact to not cross at all under the SSP1-1.9 scenario.
Datetime output
If desired, data could be interpolated first before calculating the crossing times. In such cases, returning the output as datetime rather than year might be helpful.
scmdata.processing.calculate_crossing_times(
magicc_output.resample("MS"),
threshold=2.0,
return_year=False,
)
climate_model ensemble_member model region scenario unit variable
MAGICCv7.5.1 0 unspecified World ssp585 K Surface Air Temperature Change 2042-02-01
1 unspecified World ssp585 K Surface Air Temperature Change 2041-10-01
2 unspecified World ssp585 K Surface Air Temperature Change 2035-11-01
3 unspecified World ssp585 K Surface Air Temperature Change 2035-09-01
4 unspecified World ssp585 K Surface Air Temperature Change 2036-01-01
...
595 unspecified World ssp119 K Surface Air Temperature Change NaT
596 unspecified World ssp119 K Surface Air Temperature Change NaT
597 unspecified World ssp119 K Surface Air Temperature Change NaT
598 unspecified World ssp119 K Surface Air Temperature Change 2027-07-01
599 unspecified World ssp119 K Surface Air Temperature Change NaT
Length: 4800, dtype: datetime64[ns]
Exceedance probabilities
Next we show how to calculate exceedance probabilities.
exceedance_probability_2C = scmdata.processing.calculate_exceedance_probabilities(
magicc_output,
process_over_cols=["ensemble_member", "variable"],
threshold=2.0,
)
exceedance_probability_2C
climate_model model region scenario unit
MAGICCv7.5.1 unspecified World ssp119 dimensionless 0.091667
ssp126 dimensionless 0.350000
ssp245 dimensionless 0.995000
ssp370 dimensionless 1.000000
ssp434 dimensionless 0.868333
ssp460 dimensionless 1.000000
ssp534-over dimensionless 0.983333
ssp585 dimensionless 1.000000
Name: 2.0 exceedance probability, dtype: float64
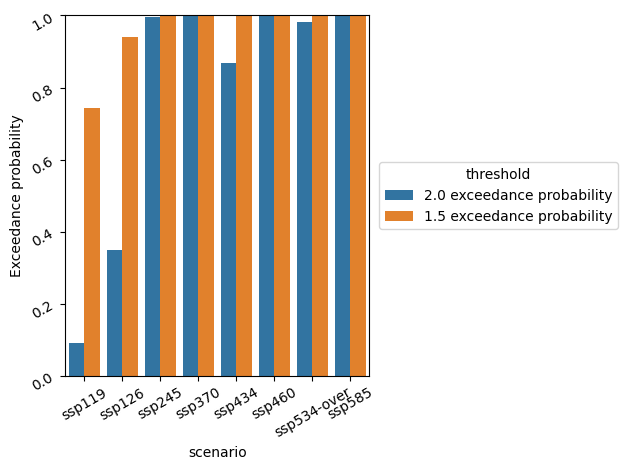
We can make a plot to compare exceedance probabilities over multiple scenarios.
exceedance_probability_15C = scmdata.processing.calculate_exceedance_probabilities(
magicc_output,
process_over_cols=["ensemble_member", "variable"],
threshold=1.5,
)
pdf = (
pd.DataFrame(
[
exceedance_probability_2C,
exceedance_probability_15C,
]
)
.T.melt(ignore_index=False, value_name="Exceedance probability")
.reset_index()
)
display(pdf) # noqa
ax = sns.barplot(data=pdf, x="scenario", y="Exceedance probability", hue="variable")
ax.tick_params(labelrotation=30)
ax.set_ylim([0, 1])
ax.legend(loc="center left", bbox_to_anchor=(1.01, 0.5), title="threshold")
plt.tight_layout()
| climate_model | model | region | scenario | unit | variable | Exceedance probability | |
|---|---|---|---|---|---|---|---|
| 0 | MAGICCv7.5.1 | unspecified | World | ssp119 | dimensionless | 2.0 exceedance probability | 0.091667 |
| 1 | MAGICCv7.5.1 | unspecified | World | ssp126 | dimensionless | 2.0 exceedance probability | 0.350000 |
| 2 | MAGICCv7.5.1 | unspecified | World | ssp245 | dimensionless | 2.0 exceedance probability | 0.995000 |
| 3 | MAGICCv7.5.1 | unspecified | World | ssp370 | dimensionless | 2.0 exceedance probability | 1.000000 |
| 4 | MAGICCv7.5.1 | unspecified | World | ssp434 | dimensionless | 2.0 exceedance probability | 0.868333 |
| 5 | MAGICCv7.5.1 | unspecified | World | ssp460 | dimensionless | 2.0 exceedance probability | 1.000000 |
| 6 | MAGICCv7.5.1 | unspecified | World | ssp534-over | dimensionless | 2.0 exceedance probability | 0.983333 |
| 7 | MAGICCv7.5.1 | unspecified | World | ssp585 | dimensionless | 2.0 exceedance probability | 1.000000 |
| 8 | MAGICCv7.5.1 | unspecified | World | ssp119 | dimensionless | 1.5 exceedance probability | 0.743333 |
| 9 | MAGICCv7.5.1 | unspecified | World | ssp126 | dimensionless | 1.5 exceedance probability | 0.940000 |
| 10 | MAGICCv7.5.1 | unspecified | World | ssp245 | dimensionless | 1.5 exceedance probability | 1.000000 |
| 11 | MAGICCv7.5.1 | unspecified | World | ssp370 | dimensionless | 1.5 exceedance probability | 1.000000 |
| 12 | MAGICCv7.5.1 | unspecified | World | ssp434 | dimensionless | 1.5 exceedance probability | 1.000000 |
| 13 | MAGICCv7.5.1 | unspecified | World | ssp460 | dimensionless | 1.5 exceedance probability | 1.000000 |
| 14 | MAGICCv7.5.1 | unspecified | World | ssp534-over | dimensionless | 1.5 exceedance probability | 1.000000 |
| 15 | MAGICCv7.5.1 | unspecified | World | ssp585 | dimensionless | 1.5 exceedance probability | 1.000000 |
Exceedance probabilities over time
It is also possible to calculate exceedance probabilities over time.
res = scmdata.processing.calculate_exceedance_probabilities_over_time(
magicc_output,
process_over_cols="ensemble_member",
threshold=1.5,
)
res = scmdata.ScmRun(res)
res.lineplot(style="variable")
<Axes: xlabel='time', ylabel='dimensionless'>
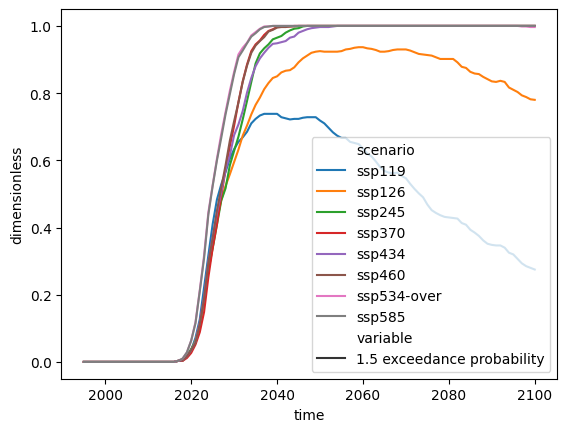
Note that taking the maximum exceedance probability over all time will be less
than or equal to the exceedance probability calculated with
calculate_exceedance_probabilities because the order of operations matters:
calculating whether each ensemble member exceeds the threshold or not then
seeing how many ensemble members out of the total exceed the threshold is not
the same as seeing how many ensemble members exceed the threshold at each
timestep and then taking the maximum over all timesteps. In general, taking
the maximum value from calculate_exceedance_probabilities_over_time will be
less than or equal to the results of calculate_exceedance_probabilities,
as demonstrated below.
comparison = (
pd.DataFrame(
{
"calculate_exceedance_probabilities": exceedance_probability_15C,
"max of calculate_exceedance_probabilities_over_time": (
res.timeseries(meta=exceedance_probability_15C.index.names).max(axis=1)
),
}
)
* 100
)
comparison.round(1)
| calculate_exceedance_probabilities | max of calculate_exceedance_probabilities_over_time | |||||
|---|---|---|---|---|---|---|
| climate_model | model | region | scenario | unit | ||
| MAGICCv7.5.1 | unspecified | World | ssp119 | dimensionless | 74.3 | 73.8 |
| ssp126 | dimensionless | 94.0 | 93.7 | |||
| ssp245 | dimensionless | 100.0 | 100.0 | |||
| ssp370 | dimensionless | 100.0 | 100.0 | |||
| ssp434 | dimensionless | 100.0 | 100.0 | |||
| ssp460 | dimensionless | 100.0 | 100.0 | |||
| ssp534-over | dimensionless | 100.0 | 100.0 | |||
| ssp585 | dimensionless | 100.0 | 100.0 |
Peak
We can calculate the peaks in each timeseries.
peak_warming = scmdata.processing.calculate_peak(magicc_output)
peak_warming
climate_model ensemble_member model region scenario unit variable
MAGICCv7.5.1 0 unspecified World ssp585 K Peak Surface Air Temperature Change 4.448044
1 unspecified World ssp585 K Peak Surface Air Temperature Change 5.372057
2 unspecified World ssp585 K Peak Surface Air Temperature Change 5.942238
3 unspecified World ssp585 K Peak Surface Air Temperature Change 5.220904
4 unspecified World ssp585 K Peak Surface Air Temperature Change 5.637467
...
595 unspecified World ssp119 K Peak Surface Air Temperature Change 1.809161
596 unspecified World ssp119 K Peak Surface Air Temperature Change 1.288471
597 unspecified World ssp119 K Peak Surface Air Temperature Change 1.669414
598 unspecified World ssp119 K Peak Surface Air Temperature Change 2.459078
599 unspecified World ssp119 K Peak Surface Air Temperature Change 1.746321
Name: 0, Length: 4800, dtype: float64
From this we can then calculate median peak warming by scenario (or other quantiles).
peak_warming.groupby(["model", "scenario"]).median()
model scenario
unspecified ssp119 1.653154
ssp126 1.878890
ssp245 2.838238
ssp370 4.372581
ssp434 2.371833
ssp460 3.485626
ssp534-over 2.613975
ssp585 5.278692
Name: 0, dtype: float64
Or make a plot.
label = "Peak warming"
pdf = peak_warming.reset_index().rename({0: label}, axis="columns")
sns.histplot(data=pdf, x=label, hue="scenario")
<Axes: xlabel='Peak warming', ylabel='Count'>
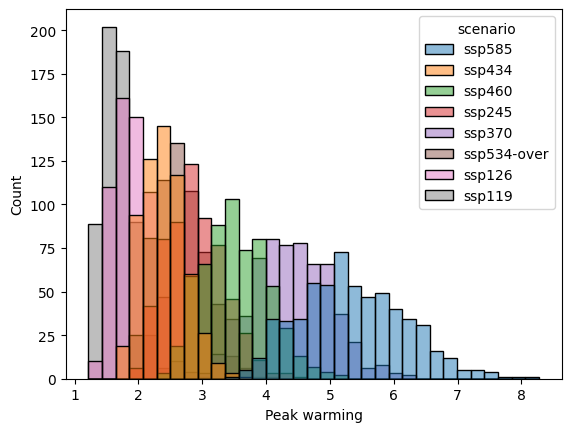
Similarly to exceedance probabilties, the order of operations matters: calculating the median of the peaks is different to calculating the median then taking the peak of this median timeseries. In general, the max of the median timeseries is less than or equal to the median of the peak in each timeseries.
comparison = pd.DataFrame(
{
"median of peak warming": peak_warming.groupby(["model", "scenario"]).median(),
"max of median timeseries": (
magicc_output.process_over(
list(set(magicc_output.meta.columns) - {"model", "scenario"}),
"median",
).max(axis=1)
),
}
)
comparison.round(3)
| median of peak warming | max of median timeseries | ||
|---|---|---|---|
| model | scenario | ||
| unspecified | ssp119 | 1.653 | 1.645 |
| ssp126 | 1.879 | 1.874 | |
| ssp245 | 2.838 | 2.838 | |
| ssp370 | 4.373 | 4.373 | |
| ssp434 | 2.372 | 2.368 | |
| ssp460 | 3.486 | 3.486 | |
| ssp534-over | 2.614 | 2.613 | |
| ssp585 | 5.279 | 5.279 |
Peak time
We can calculate the peak time in each timeseries.
peak_warming_year = scmdata.processing.calculate_peak_time(magicc_output)
peak_warming_year
climate_model ensemble_member model region scenario unit variable
MAGICCv7.5.1 0 unspecified World ssp585 K Year of peak Surface Air Temperature Change 2100
1 unspecified World ssp585 K Year of peak Surface Air Temperature Change 2100
2 unspecified World ssp585 K Year of peak Surface Air Temperature Change 2100
3 unspecified World ssp585 K Year of peak Surface Air Temperature Change 2100
4 unspecified World ssp585 K Year of peak Surface Air Temperature Change 2100
...
595 unspecified World ssp119 K Year of peak Surface Air Temperature Change 2048
596 unspecified World ssp119 K Year of peak Surface Air Temperature Change 2036
597 unspecified World ssp119 K Year of peak Surface Air Temperature Change 2039
598 unspecified World ssp119 K Year of peak Surface Air Temperature Change 2059
599 unspecified World ssp119 K Year of peak Surface Air Temperature Change 2048
Name: 0, Length: 4800, dtype: int64
From this we can then calculate median peak warming time by scenario (or other quantiles).
peak_warming_year.groupby(["model", "scenario"]).median()
model scenario
unspecified ssp119 2048.0
ssp126 2069.0
ssp245 2100.0
ssp370 2100.0
ssp434 2094.0
ssp460 2100.0
ssp534-over 2060.0
ssp585 2100.0
Name: 0, dtype: float64
label = "Peak warming year"
pdf = peak_warming_year.reset_index().rename({0: label}, axis="columns")
ax = sns.histplot(data=pdf, x=label, hue="scenario", bins=np.arange(2025, 2100 + 1))
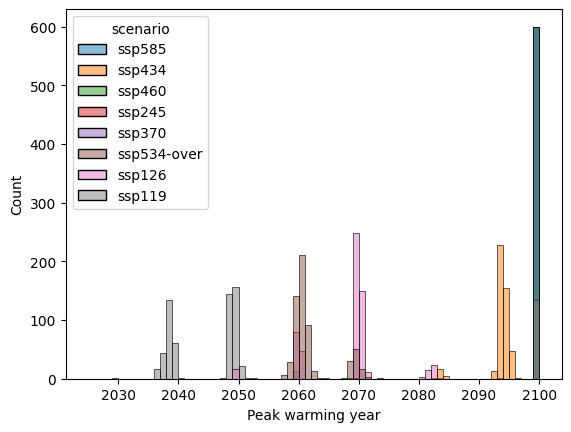
SR1.5 categorisation
It is also possible to categorise the scenarios using the same categorisation as in SR1.5. To do this we have to first calculate the appropriate quantiles.
magicc_output_categorisation_quantiles = scmdata.ScmRun(
magicc_output.quantiles_over("ensemble_member", quantiles=[0.33, 0.5, 0.66])
)
magicc_output_categorisation_quantiles
/home/docs/checkouts/readthedocs.org/user_builds/scmdata/checkouts/stable/src/scmdata/run.py:197: PerformanceWarning: DataFrame is highly fragmented. This is usually the result of calling `frame.insert` many times, which has poor performance. Consider joining all columns at once using pd.concat(axis=1) instead. To get a de-fragmented frame, use `newframe = frame.copy()`
df.reset_index(inplace=True)
/home/docs/checkouts/readthedocs.org/user_builds/scmdata/checkouts/stable/src/scmdata/run.py:197: PerformanceWarning: DataFrame is highly fragmented. This is usually the result of calling `frame.insert` many times, which has poor performance. Consider joining all columns at once using pd.concat(axis=1) instead. To get a de-fragmented frame, use `newframe = frame.copy()`
df.reset_index(inplace=True)
/home/docs/checkouts/readthedocs.org/user_builds/scmdata/checkouts/stable/src/scmdata/run.py:197: PerformanceWarning: DataFrame is highly fragmented. This is usually the result of calling `frame.insert` many times, which has poor performance. Consider joining all columns at once using pd.concat(axis=1) instead. To get a de-fragmented frame, use `newframe = frame.copy()`
df.reset_index(inplace=True)
/home/docs/checkouts/readthedocs.org/user_builds/scmdata/checkouts/stable/src/scmdata/run.py:197: PerformanceWarning: DataFrame is highly fragmented. This is usually the result of calling `frame.insert` many times, which has poor performance. Consider joining all columns at once using pd.concat(axis=1) instead. To get a de-fragmented frame, use `newframe = frame.copy()`
df.reset_index(inplace=True)
/home/docs/checkouts/readthedocs.org/user_builds/scmdata/checkouts/stable/src/scmdata/run.py:197: PerformanceWarning: DataFrame is highly fragmented. This is usually the result of calling `frame.insert` many times, which has poor performance. Consider joining all columns at once using pd.concat(axis=1) instead. To get a de-fragmented frame, use `newframe = frame.copy()`
df.reset_index(inplace=True)
/home/docs/checkouts/readthedocs.org/user_builds/scmdata/checkouts/stable/src/scmdata/run.py:197: PerformanceWarning: DataFrame is highly fragmented. This is usually the result of calling `frame.insert` many times, which has poor performance. Consider joining all columns at once using pd.concat(axis=1) instead. To get a de-fragmented frame, use `newframe = frame.copy()`
df.reset_index(inplace=True)
/home/docs/checkouts/readthedocs.org/user_builds/scmdata/checkouts/stable/src/scmdata/run.py:197: PerformanceWarning: DataFrame is highly fragmented. This is usually the result of calling `frame.insert` many times, which has poor performance. Consider joining all columns at once using pd.concat(axis=1) instead. To get a de-fragmented frame, use `newframe = frame.copy()`
df.reset_index(inplace=True)
<ScmRun (timeseries: 24, timepoints: 106)>
Time:
Start: 1995-01-01T00:00:00
End: 2100-01-01T00:00:00
Meta:
climate_model model quantile region scenario unit \
0 MAGICCv7.5.1 unspecified 0.33 World ssp119 K
1 MAGICCv7.5.1 unspecified 0.33 World ssp126 K
2 MAGICCv7.5.1 unspecified 0.33 World ssp245 K
3 MAGICCv7.5.1 unspecified 0.33 World ssp370 K
4 MAGICCv7.5.1 unspecified 0.33 World ssp434 K
5 MAGICCv7.5.1 unspecified 0.33 World ssp460 K
6 MAGICCv7.5.1 unspecified 0.33 World ssp534-over K
7 MAGICCv7.5.1 unspecified 0.33 World ssp585 K
8 MAGICCv7.5.1 unspecified 0.50 World ssp119 K
9 MAGICCv7.5.1 unspecified 0.50 World ssp126 K
10 MAGICCv7.5.1 unspecified 0.50 World ssp245 K
11 MAGICCv7.5.1 unspecified 0.50 World ssp370 K
12 MAGICCv7.5.1 unspecified 0.50 World ssp434 K
13 MAGICCv7.5.1 unspecified 0.50 World ssp460 K
14 MAGICCv7.5.1 unspecified 0.50 World ssp534-over K
15 MAGICCv7.5.1 unspecified 0.50 World ssp585 K
16 MAGICCv7.5.1 unspecified 0.66 World ssp119 K
17 MAGICCv7.5.1 unspecified 0.66 World ssp126 K
18 MAGICCv7.5.1 unspecified 0.66 World ssp245 K
19 MAGICCv7.5.1 unspecified 0.66 World ssp370 K
20 MAGICCv7.5.1 unspecified 0.66 World ssp434 K
21 MAGICCv7.5.1 unspecified 0.66 World ssp460 K
22 MAGICCv7.5.1 unspecified 0.66 World ssp534-over K
23 MAGICCv7.5.1 unspecified 0.66 World ssp585 K
variable
0 Surface Air Temperature Change
1 Surface Air Temperature Change
2 Surface Air Temperature Change
3 Surface Air Temperature Change
4 Surface Air Temperature Change
5 Surface Air Temperature Change
6 Surface Air Temperature Change
7 Surface Air Temperature Change
8 Surface Air Temperature Change
9 Surface Air Temperature Change
10 Surface Air Temperature Change
11 Surface Air Temperature Change
12 Surface Air Temperature Change
13 Surface Air Temperature Change
14 Surface Air Temperature Change
15 Surface Air Temperature Change
16 Surface Air Temperature Change
17 Surface Air Temperature Change
18 Surface Air Temperature Change
19 Surface Air Temperature Change
20 Surface Air Temperature Change
21 Surface Air Temperature Change
22 Surface Air Temperature Change
23 Surface Air Temperature Change
scmdata.processing.categorisation_sr15(
magicc_output_categorisation_quantiles, ["climate_model", "scenario"]
)
climate_model scenario
MAGICCv7.5.1 ssp119 1.5C high overshoot
ssp126 Higher 2C
ssp245 Above 2C
ssp370 Above 2C
ssp434 Above 2C
ssp460 Above 2C
ssp534-over Above 2C
ssp585 Above 2C
Name: category, dtype: object
Set of summary variables
It is also possible to calculate a set of summary variables using the convenience function
calculate_summary_stats. The documentation is given below.
print(scmdata.processing.calculate_summary_stats.__doc__)
Calculate common summary statistics
Parameters
----------
scmrun : :class:`scmdata.ScmRun`
Data of which to calculate the stats
index : list[str]
Columns to use in the index of the output (unit is added if not
included)
exceedance_probabilities_thresholds : list[float]
Thresholds to use for exceedance probabilities
exceedance_probabilities_variable : str
Variable to use for exceedance probability calculations
exceedance_probabilities_naming_base : str
String to use as the base for naming the exceedance probabilities. Each
exceedance probability output column will have a name given by
``exceedance_probabilities_naming_base.format(threshold)`` where
threshold is the exceedance probability threshold to use. If not
supplied, the default output of
:func:`scmdata.processing.calculate_exceedance_probabilities` will be
used.
peak_quantiles : list[float]
Quantiles to report in peak calculations
peak_variable : str
Variable of which to calculate the peak
peak_naming_base : str
Base to use for naming the peak outputs. This is combined with the
quantile. If not supplied, ``"{} peak"`` is used so the outputs will be
named e.g. "0.05 peak", "0.5 peak", "0.95 peak".
peak_time_naming_base : str
Base to use for naming the peak time outputs. This is combined with the
quantile. If not supplied, ``"{} peak year"`` is used (unless
``peak_return_year`` is ``False`` in which case ``"{} peak time"`` is
used) so the outputs will be named e.g. "0.05 peak year", "0.5 peak
year", "0.95 peak year".
peak_return_year : bool
If ``True``, return the year of the peak of ``peak_variable``,
otherwise return full dates
categorisation_variable : str
Variable to use for categorisation. Note that this variable point to
timeseries that contain global-mean surface air temperatures (GSAT)
relative to 1850-1900 (using another reference period will not break
this function, but is inconsistent with the original algorithm).
categorisation_quantile_cols : list[str]
Columns which represent individual ensemble members in the output (e.g.
["ensemble_member"]). The quantiles are taking over these columns
before the data is passed to
:func:`scmdata.processing.categorisation_sr15`.
progress : bool
Should a progress bar be shown whilst the calculations are done?
Returns
-------
:class:`pd.DataFrame`
Summary statistics, with each column being a statistic and the index
being given by ``index``
It can be used to calculate summary statistics as shown below.
summary_stats = scmdata.processing.calculate_summary_stats(
magicc_output,
["climate_model", "model", "scenario", "region"],
exceedance_probabilities_thresholds=np.arange(1, 2.51, 0.1),
exceedance_probabilities_variable="Surface Air Temperature Change",
exceedance_probabilities_naming_base="Exceedance Probability|{:.2f}K",
peak_quantiles=[0.05, 0.5, 0.95],
peak_variable="Surface Air Temperature Change",
peak_naming_base="Peak Surface Air Temperature Change|{}th quantile",
peak_time_naming_base="Year of peak Surface Air Temperature Change|{}th quantile",
peak_return_year=True,
progress=True,
)
summary_stats
/home/docs/checkouts/readthedocs.org/user_builds/scmdata/checkouts/stable/src/scmdata/run.py:197: PerformanceWarning: DataFrame is highly fragmented. This is usually the result of calling `frame.insert` many times, which has poor performance. Consider joining all columns at once using pd.concat(axis=1) instead. To get a de-fragmented frame, use `newframe = frame.copy()`
df.reset_index(inplace=True)
/home/docs/checkouts/readthedocs.org/user_builds/scmdata/checkouts/stable/src/scmdata/run.py:197: PerformanceWarning: DataFrame is highly fragmented. This is usually the result of calling `frame.insert` many times, which has poor performance. Consider joining all columns at once using pd.concat(axis=1) instead. To get a de-fragmented frame, use `newframe = frame.copy()`
df.reset_index(inplace=True)
/home/docs/checkouts/readthedocs.org/user_builds/scmdata/checkouts/stable/src/scmdata/run.py:197: PerformanceWarning: DataFrame is highly fragmented. This is usually the result of calling `frame.insert` many times, which has poor performance. Consider joining all columns at once using pd.concat(axis=1) instead. To get a de-fragmented frame, use `newframe = frame.copy()`
df.reset_index(inplace=True)
/home/docs/checkouts/readthedocs.org/user_builds/scmdata/checkouts/stable/src/scmdata/run.py:197: PerformanceWarning: DataFrame is highly fragmented. This is usually the result of calling `frame.insert` many times, which has poor performance. Consider joining all columns at once using pd.concat(axis=1) instead. To get a de-fragmented frame, use `newframe = frame.copy()`
df.reset_index(inplace=True)
/home/docs/checkouts/readthedocs.org/user_builds/scmdata/checkouts/stable/src/scmdata/run.py:197: PerformanceWarning: DataFrame is highly fragmented. This is usually the result of calling `frame.insert` many times, which has poor performance. Consider joining all columns at once using pd.concat(axis=1) instead. To get a de-fragmented frame, use `newframe = frame.copy()`
df.reset_index(inplace=True)
/home/docs/checkouts/readthedocs.org/user_builds/scmdata/checkouts/stable/src/scmdata/run.py:197: PerformanceWarning: DataFrame is highly fragmented. This is usually the result of calling `frame.insert` many times, which has poor performance. Consider joining all columns at once using pd.concat(axis=1) instead. To get a de-fragmented frame, use `newframe = frame.copy()`
df.reset_index(inplace=True)
/home/docs/checkouts/readthedocs.org/user_builds/scmdata/checkouts/stable/src/scmdata/run.py:197: PerformanceWarning: DataFrame is highly fragmented. This is usually the result of calling `frame.insert` many times, which has poor performance. Consider joining all columns at once using pd.concat(axis=1) instead. To get a de-fragmented frame, use `newframe = frame.copy()`
df.reset_index(inplace=True)
0%| | 0/23 [00:00<?, ?it/s]
22%|██▏ | 5/23 [00:00<00:00, 49.64it/s]
48%|████▊ | 11/23 [00:00<00:00, 50.38it/s]
74%|███████▍ | 17/23 [00:00<00:00, 53.27it/s]
100%|██████████| 23/23 [00:00<00:00, 54.97it/s]
100%|██████████| 23/23 [00:00<00:00, 53.52it/s]
climate_model model scenario region unit statistic
MAGICCv7.5.1 unspecified ssp119 World dimensionless Exceedance Probability|1.00K 1.0
Exceedance Probability|1.10K 1.0
Exceedance Probability|1.20K 1.0
Exceedance Probability|1.30K 0.976667
Exceedance Probability|1.40K 0.888333
...
ssp370 World SR1.5 category Above 2C
ssp434 World SR1.5 category Above 2C
ssp460 World SR1.5 category Above 2C
ssp534-over World SR1.5 category Above 2C
ssp585 World SR1.5 category Above 2C
Name: value, Length: 184, dtype: object
We can then use pandas to create summary tables of interest.
summary_stats.unstack(["climate_model", "statistic", "unit"])
| climate_model | MAGICCv7.5.1 | ||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| statistic | Exceedance Probability|1.00K | Exceedance Probability|1.10K | Exceedance Probability|1.20K | Exceedance Probability|1.30K | Exceedance Probability|1.40K | Exceedance Probability|1.50K | Exceedance Probability|1.60K | Exceedance Probability|1.70K | Exceedance Probability|1.80K | Exceedance Probability|1.90K | ... | Exceedance Probability|2.30K | Exceedance Probability|2.40K | Exceedance Probability|2.50K | Peak Surface Air Temperature Change|0.05th quantile | Peak Surface Air Temperature Change|0.5th quantile | Peak Surface Air Temperature Change|0.95th quantile | Year of peak Surface Air Temperature Change|0.05th quantile | Year of peak Surface Air Temperature Change|0.5th quantile | Year of peak Surface Air Temperature Change|0.95th quantile | SR1.5 category | ||
| unit | dimensionless | dimensionless | dimensionless | dimensionless | dimensionless | dimensionless | dimensionless | dimensionless | dimensionless | dimensionless | ... | dimensionless | dimensionless | dimensionless | K | K | K | K | K | K | |||
| model | scenario | region | |||||||||||||||||||||
| unspecified | ssp119 | World | 1.0 | 1.0 | 1.0 | 0.976667 | 0.888333 | 0.743333 | 0.608333 | 0.418333 | 0.286667 | 0.156667 | ... | 0.008333 | 0.003333 | 0.0 | 1.345672 | 1.653154 | 2.071348 | 2037.0 | 2048.0 | 2050.0 | 1.5C high overshoot |
| ssp126 | World | 1.0 | 1.0 | 1.0 | 1.0 | 0.988333 | 0.94 | 0.836667 | 0.735 | 0.618333 | 0.465 | ... | 0.096667 | 0.056667 | 0.025 | 1.478604 | 1.87889 | 2.425129 | 2059.0 | 2069.0 | 2081.0 | Higher 2C | |
| ssp245 | World | 1.0 | 1.0 | 1.0 | 1.0 | 1.0 | 1.0 | 1.0 | 1.0 | 1.0 | 1.0 | ... | 0.911667 | 0.846667 | 0.783333 | 2.218657 | 2.838238 | 3.642782 | 2100.0 | 2100.0 | 2100.0 | Above 2C | |
| ssp370 | World | 1.0 | 1.0 | 1.0 | 1.0 | 1.0 | 1.0 | 1.0 | 1.0 | 1.0 | 1.0 | ... | 1.0 | 1.0 | 1.0 | 3.476406 | 4.372581 | 5.35643 | 2100.0 | 2100.0 | 2100.0 | Above 2C | |
| ssp434 | World | 1.0 | 1.0 | 1.0 | 1.0 | 1.0 | 1.0 | 1.0 | 0.996667 | 0.978333 | 0.946667 | ... | 0.58 | 0.463333 | 0.356667 | 1.898714 | 2.371833 | 2.958278 | 2092.0 | 2094.0 | 2100.0 | Above 2C | |
| ssp460 | World | 1.0 | 1.0 | 1.0 | 1.0 | 1.0 | 1.0 | 1.0 | 1.0 | 1.0 | 1.0 | ... | 1.0 | 1.0 | 0.995 | 2.753816 | 3.485626 | 4.373157 | 2100.0 | 2100.0 | 2100.0 | Above 2C | |
| ssp534-over | World | 1.0 | 1.0 | 1.0 | 1.0 | 1.0 | 1.0 | 1.0 | 1.0 | 1.0 | 0.996667 | ... | 0.811667 | 0.725 | 0.63 | 2.100912 | 2.613975 | 3.299928 | 2058.0 | 2060.0 | 2069.0 | Above 2C | |
| ssp585 | World | 1.0 | 1.0 | 1.0 | 1.0 | 1.0 | 1.0 | 1.0 | 1.0 | 1.0 | 1.0 | ... | 1.0 | 1.0 | 1.0 | 4.094323 | 5.278692 | 6.75008 | 2100.0 | 2100.0 | 2100.0 | Above 2C | |
8 rows × 23 columns
index = ["climate_model", "scenario"]
pivot_merge_unit = summary_stats.to_frame().reset_index()
pivot_merge_unit["statistic"] = pivot_merge_unit["statistic"] + pivot_merge_unit[
"unit"
].apply(lambda x: f"({x})" if x else "")
pivot_merge_unit = pivot_merge_unit.drop("unit", axis="columns")
pivot_merge_unit = pivot_merge_unit.set_index(
list(set(pivot_merge_unit.columns) - {"value"})
).unstack("statistic")
pivot_merge_unit
| value | ||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| statistic | Exceedance Probability|1.00K(dimensionless) | Exceedance Probability|1.10K(dimensionless) | Exceedance Probability|1.20K(dimensionless) | Exceedance Probability|1.30K(dimensionless) | Exceedance Probability|1.40K(dimensionless) | Exceedance Probability|1.50K(dimensionless) | Exceedance Probability|1.60K(dimensionless) | Exceedance Probability|1.70K(dimensionless) | Exceedance Probability|1.80K(dimensionless) | Exceedance Probability|1.90K(dimensionless) | ... | Exceedance Probability|2.30K(dimensionless) | Exceedance Probability|2.40K(dimensionless) | Exceedance Probability|2.50K(dimensionless) | Peak Surface Air Temperature Change|0.05th quantile(K) | Peak Surface Air Temperature Change|0.5th quantile(K) | Peak Surface Air Temperature Change|0.95th quantile(K) | SR1.5 category | Year of peak Surface Air Temperature Change|0.05th quantile(K) | Year of peak Surface Air Temperature Change|0.5th quantile(K) | Year of peak Surface Air Temperature Change|0.95th quantile(K) | |||
| region | scenario | model | climate_model | |||||||||||||||||||||
| World | ssp119 | unspecified | MAGICCv7.5.1 | 1.0 | 1.0 | 1.0 | 0.976667 | 0.888333 | 0.743333 | 0.608333 | 0.418333 | 0.286667 | 0.156667 | ... | 0.008333 | 0.003333 | 0.0 | 1.345672 | 1.653154 | 2.071348 | 1.5C high overshoot | 2037.0 | 2048.0 | 2050.0 |
| ssp126 | unspecified | MAGICCv7.5.1 | 1.0 | 1.0 | 1.0 | 1.0 | 0.988333 | 0.94 | 0.836667 | 0.735 | 0.618333 | 0.465 | ... | 0.096667 | 0.056667 | 0.025 | 1.478604 | 1.87889 | 2.425129 | Higher 2C | 2059.0 | 2069.0 | 2081.0 | |
| ssp245 | unspecified | MAGICCv7.5.1 | 1.0 | 1.0 | 1.0 | 1.0 | 1.0 | 1.0 | 1.0 | 1.0 | 1.0 | 1.0 | ... | 0.911667 | 0.846667 | 0.783333 | 2.218657 | 2.838238 | 3.642782 | Above 2C | 2100.0 | 2100.0 | 2100.0 | |
| ssp370 | unspecified | MAGICCv7.5.1 | 1.0 | 1.0 | 1.0 | 1.0 | 1.0 | 1.0 | 1.0 | 1.0 | 1.0 | 1.0 | ... | 1.0 | 1.0 | 1.0 | 3.476406 | 4.372581 | 5.35643 | Above 2C | 2100.0 | 2100.0 | 2100.0 | |
| ssp434 | unspecified | MAGICCv7.5.1 | 1.0 | 1.0 | 1.0 | 1.0 | 1.0 | 1.0 | 1.0 | 0.996667 | 0.978333 | 0.946667 | ... | 0.58 | 0.463333 | 0.356667 | 1.898714 | 2.371833 | 2.958278 | Above 2C | 2092.0 | 2094.0 | 2100.0 | |
| ssp460 | unspecified | MAGICCv7.5.1 | 1.0 | 1.0 | 1.0 | 1.0 | 1.0 | 1.0 | 1.0 | 1.0 | 1.0 | 1.0 | ... | 1.0 | 1.0 | 0.995 | 2.753816 | 3.485626 | 4.373157 | Above 2C | 2100.0 | 2100.0 | 2100.0 | |
| ssp534-over | unspecified | MAGICCv7.5.1 | 1.0 | 1.0 | 1.0 | 1.0 | 1.0 | 1.0 | 1.0 | 1.0 | 1.0 | 0.996667 | ... | 0.811667 | 0.725 | 0.63 | 2.100912 | 2.613975 | 3.299928 | Above 2C | 2058.0 | 2060.0 | 2069.0 | |
| ssp585 | unspecified | MAGICCv7.5.1 | 1.0 | 1.0 | 1.0 | 1.0 | 1.0 | 1.0 | 1.0 | 1.0 | 1.0 | 1.0 | ... | 1.0 | 1.0 | 1.0 | 4.094323 | 5.278692 | 6.75008 | Above 2C | 2100.0 | 2100.0 | 2100.0 | |
8 rows × 23 columns