import pathlib
NetCDF handling
NetCDF formatted files are much faster to read and write for large datasets.
In order to make the most of this, the ScmRun objects have the ability to
read and write netCDF files.
import traceback
from tempfile import TemporaryDirectory
import numpy as np
import seaborn as sns
import xarray as xr
from scmdata.netcdf import nc_to_run
from scmdata.run import ScmRun, run_append
/home/docs/checkouts/readthedocs.org/user_builds/scmdata/checkouts/stable/src/scmdata/database/_database.py:9: TqdmWarning: IProgress not found. Please update jupyter and ipywidgets. See https://ipywidgets.readthedocs.io/en/stable/user_install.html
import tqdm.autonotebook as tqdman
Helper bits and piecs
temp_directory = TemporaryDirectory()
generator = np.random.default_rng(0)
OUTPUT_DIR = pathlib.Path(temp_directory.name)
OUT_FNAME = OUTPUT_DIR / "out_runs.nc"
def new_timeseries( # noqa: PLR0913
n=100,
count=1,
model="example",
scenario="ssp119",
variable="Surface Temperature",
unit="K",
region="World",
cls=ScmRun,
**kwargs,
):
"""
Create an example timeseries
"""
data = generator.random((n, count)) * np.arange(n)[:, np.newaxis]
index = 2000 + np.arange(n)
return cls(
data,
columns={
"model": model,
"scenario": scenario,
"variable": variable,
"region": region,
"unit": unit,
**kwargs,
},
index=index,
)
Let’s create an ScmRun which contains a few variables and a number of runs.
Such a dataframe would be used to store the results from an ensemble of
simple climate model runs.
runs = run_append(
[
new_timeseries(
count=3,
variable=[
"Surface Temperature",
"Atmospheric Concentrations|CO2",
"Radiative Forcing",
],
unit=["K", "ppm", "W/m^2"],
run_id=run_id,
)
for run_id in range(10)
]
)
runs.metadata["source"] = "fake data"
runs
<ScmRun (timeseries: 30, timepoints: 100)>
Time:
Start: 2000-01-01T00:00:00
End: 2099-01-01T00:00:00
Meta:
model region run_id scenario unit variable
0 example World 0 ssp119 K Surface Temperature
1 example World 0 ssp119 ppm Atmospheric Concentrations|CO2
2 example World 0 ssp119 W/m^2 Radiative Forcing
3 example World 1 ssp119 K Surface Temperature
4 example World 1 ssp119 ppm Atmospheric Concentrations|CO2
5 example World 1 ssp119 W/m^2 Radiative Forcing
6 example World 2 ssp119 K Surface Temperature
7 example World 2 ssp119 ppm Atmospheric Concentrations|CO2
8 example World 2 ssp119 W/m^2 Radiative Forcing
9 example World 3 ssp119 K Surface Temperature
10 example World 3 ssp119 ppm Atmospheric Concentrations|CO2
11 example World 3 ssp119 W/m^2 Radiative Forcing
12 example World 4 ssp119 K Surface Temperature
13 example World 4 ssp119 ppm Atmospheric Concentrations|CO2
14 example World 4 ssp119 W/m^2 Radiative Forcing
15 example World 5 ssp119 K Surface Temperature
16 example World 5 ssp119 ppm Atmospheric Concentrations|CO2
17 example World 5 ssp119 W/m^2 Radiative Forcing
18 example World 6 ssp119 K Surface Temperature
19 example World 6 ssp119 ppm Atmospheric Concentrations|CO2
20 example World 6 ssp119 W/m^2 Radiative Forcing
21 example World 7 ssp119 K Surface Temperature
22 example World 7 ssp119 ppm Atmospheric Concentrations|CO2
23 example World 7 ssp119 W/m^2 Radiative Forcing
24 example World 8 ssp119 K Surface Temperature
25 example World 8 ssp119 ppm Atmospheric Concentrations|CO2
26 example World 8 ssp119 W/m^2 Radiative Forcing
27 example World 9 ssp119 K Surface Temperature
28 example World 9 ssp119 ppm Atmospheric Concentrations|CO2
29 example World 9 ssp119 W/m^2 Radiative Forcing
Reading/Writing to NetCDF4
Basics
Writing the runs to disk is easy. The one trick is that each variable and dimension combination must have unique metadata. If they do not, you will receive an error message like the below.
try:
runs.to_nc(OUT_FNAME, dimensions=["region"])
except ValueError:
traceback.print_exc(limit=0, chain=False)
ValueError: dimensions: `['region']` and extras: `[]` do not uniquely define the timeseries, please add extra dimensions and/or extras
In our dataset, there is more than one “run_id” per variable hence we need to use a different
dimension, run_id, because this will result in each variable’s remaining metadata being unique.
runs.to_nc(OUT_FNAME, dimensions=["run_id"])
/home/docs/checkouts/readthedocs.org/user_builds/scmdata/checkouts/stable/src/scmdata/_xarray.py:236: FutureWarning: The previous implementation of stack is deprecated and will be removed in a future version of pandas. See the What's New notes for pandas 2.1.0 for details. Specify future_stack=True to adopt the new implementation and silence this warning.
else timeseries.T.stack(dimensions)
The output netCDF file can be read using the from_nc method, nc_to_run function or directly
using xarray.
runs_netcdf = ScmRun.from_nc(OUT_FNAME)
runs_netcdf
<ScmRun (timeseries: 30, timepoints: 100)>
Time:
Start: 2000-01-01T00:00:00
End: 2099-01-01T00:00:00
Meta:
model region run_id scenario unit variable
0 example World 0 ssp119 K Surface Temperature
1 example World 0 ssp119 ppm Atmospheric Concentrations|CO2
2 example World 0 ssp119 W/m^2 Radiative Forcing
3 example World 1 ssp119 K Surface Temperature
4 example World 1 ssp119 ppm Atmospheric Concentrations|CO2
5 example World 1 ssp119 W/m^2 Radiative Forcing
6 example World 2 ssp119 K Surface Temperature
7 example World 2 ssp119 ppm Atmospheric Concentrations|CO2
8 example World 2 ssp119 W/m^2 Radiative Forcing
9 example World 3 ssp119 K Surface Temperature
10 example World 3 ssp119 ppm Atmospheric Concentrations|CO2
11 example World 3 ssp119 W/m^2 Radiative Forcing
12 example World 4 ssp119 K Surface Temperature
13 example World 4 ssp119 ppm Atmospheric Concentrations|CO2
14 example World 4 ssp119 W/m^2 Radiative Forcing
15 example World 5 ssp119 K Surface Temperature
16 example World 5 ssp119 ppm Atmospheric Concentrations|CO2
17 example World 5 ssp119 W/m^2 Radiative Forcing
18 example World 6 ssp119 K Surface Temperature
19 example World 6 ssp119 ppm Atmospheric Concentrations|CO2
20 example World 6 ssp119 W/m^2 Radiative Forcing
21 example World 7 ssp119 K Surface Temperature
22 example World 7 ssp119 ppm Atmospheric Concentrations|CO2
23 example World 7 ssp119 W/m^2 Radiative Forcing
24 example World 8 ssp119 K Surface Temperature
25 example World 8 ssp119 ppm Atmospheric Concentrations|CO2
26 example World 8 ssp119 W/m^2 Radiative Forcing
27 example World 9 ssp119 K Surface Temperature
28 example World 9 ssp119 ppm Atmospheric Concentrations|CO2
29 example World 9 ssp119 W/m^2 Radiative Forcing
nc_to_run(ScmRun, OUT_FNAME)
<ScmRun (timeseries: 30, timepoints: 100)>
Time:
Start: 2000-01-01T00:00:00
End: 2099-01-01T00:00:00
Meta:
model region run_id scenario unit variable
0 example World 0 ssp119 K Surface Temperature
1 example World 0 ssp119 ppm Atmospheric Concentrations|CO2
2 example World 0 ssp119 W/m^2 Radiative Forcing
3 example World 1 ssp119 K Surface Temperature
4 example World 1 ssp119 ppm Atmospheric Concentrations|CO2
5 example World 1 ssp119 W/m^2 Radiative Forcing
6 example World 2 ssp119 K Surface Temperature
7 example World 2 ssp119 ppm Atmospheric Concentrations|CO2
8 example World 2 ssp119 W/m^2 Radiative Forcing
9 example World 3 ssp119 K Surface Temperature
10 example World 3 ssp119 ppm Atmospheric Concentrations|CO2
11 example World 3 ssp119 W/m^2 Radiative Forcing
12 example World 4 ssp119 K Surface Temperature
13 example World 4 ssp119 ppm Atmospheric Concentrations|CO2
14 example World 4 ssp119 W/m^2 Radiative Forcing
15 example World 5 ssp119 K Surface Temperature
16 example World 5 ssp119 ppm Atmospheric Concentrations|CO2
17 example World 5 ssp119 W/m^2 Radiative Forcing
18 example World 6 ssp119 K Surface Temperature
19 example World 6 ssp119 ppm Atmospheric Concentrations|CO2
20 example World 6 ssp119 W/m^2 Radiative Forcing
21 example World 7 ssp119 K Surface Temperature
22 example World 7 ssp119 ppm Atmospheric Concentrations|CO2
23 example World 7 ssp119 W/m^2 Radiative Forcing
24 example World 8 ssp119 K Surface Temperature
25 example World 8 ssp119 ppm Atmospheric Concentrations|CO2
26 example World 8 ssp119 W/m^2 Radiative Forcing
27 example World 9 ssp119 K Surface Temperature
28 example World 9 ssp119 ppm Atmospheric Concentrations|CO2
29 example World 9 ssp119 W/m^2 Radiative Forcing
xr.load_dataset(OUT_FNAME)
<xarray.Dataset>
Dimensions: (time: 100, run_id: 10)
Coordinates:
* time (time) datetime64[ns] 2000-01-01 ... 209...
* run_id (run_id) int64 0 1 2 3 4 5 6 7 8 9
Data variables:
Surface_Temperature (run_id, time) float64 0.0 ... 2.459
Atmospheric_Concentrations__CO2 (run_id, time) float64 0.0 0.8133 ... 20.56
Radiative_Forcing (run_id, time) float64 0.0 0.9128 ... 29.75
Attributes:
scmdata_metadata_scenario: ssp119
scmdata_metadata_model: example
scmdata_metadata_region: World
created_at: 2024-01-29T07:18:01.855754
_scmdata_version: 1.0.0
source: fake dataThe additional metadata in runs is also serialized and deserialized in the netCDF files. The
metadata of the loaded ScmRun will also contain some additional fields about the file
creation.
assert "source" in runs_netcdf.metadata
runs_netcdf.metadata
{'created_at': '2024-01-29T07:18:01.855754',
'_scmdata_version': '1.0.0',
'source': 'fake data'}
Splitting your data
Sometimes if you have complicated ensemble runs it might be more efficient to split the data into smaller subsets.
In the below example we iterate over scenarios to produce a netCDF file per scenario.
large_run = []
# 10 runs for each scenario
for sce in ["ssp119", "ssp370", "ssp585"]:
large_run.extend(
[
new_timeseries(
count=3,
scenario=sce,
variable=[
"Surface Temperature",
"Atmospheric Concentrations|CO2",
"Radiative Forcing",
],
unit=["K", "ppm", "W/m^2"],
paraset_id=paraset_id,
)
for paraset_id in range(10)
]
)
large_run = run_append(large_run)
# also set a run_id (often we'd have paraset_id and run_id,
# one which keeps track of the parameter set we've run and
# the other which keeps track of the run in a large ensemble)
large_run["run_id"] = large_run.meta.index.values
large_run
<ScmRun (timeseries: 90, timepoints: 100)>
Time:
Start: 2000-01-01T00:00:00
End: 2099-01-01T00:00:00
Meta:
model paraset_id region run_id scenario unit \
0 example 0 World 0 ssp119 K
1 example 0 World 1 ssp119 ppm
2 example 0 World 2 ssp119 W/m^2
3 example 1 World 3 ssp119 K
4 example 1 World 4 ssp119 ppm
.. ... ... ... ... ... ...
85 example 8 World 85 ssp585 ppm
86 example 8 World 86 ssp585 W/m^2
87 example 9 World 87 ssp585 K
88 example 9 World 88 ssp585 ppm
89 example 9 World 89 ssp585 W/m^2
variable
0 Surface Temperature
1 Atmospheric Concentrations|CO2
2 Radiative Forcing
3 Surface Temperature
4 Atmospheric Concentrations|CO2
.. ...
85 Atmospheric Concentrations|CO2
86 Radiative Forcing
87 Surface Temperature
88 Atmospheric Concentrations|CO2
89 Radiative Forcing
[90 rows x 7 columns]
Data for each scenario can then be loaded independently instead of having to load all the data and then filtering
for sce_run in large_run.groupby("scenario"):
sce = sce_run.get_unique_meta("scenario", True)
sce_run.to_nc(
OUTPUT_DIR / f"out-{sce}-sparse.nc",
dimensions=["run_id", "paraset_id"],
)
/home/docs/checkouts/readthedocs.org/user_builds/scmdata/checkouts/stable/src/scmdata/_xarray.py:236: FutureWarning: The previous implementation of stack is deprecated and will be removed in a future version of pandas. See the What's New notes for pandas 2.1.0 for details. Specify future_stack=True to adopt the new implementation and silence this warning.
else timeseries.T.stack(dimensions)
/home/docs/checkouts/readthedocs.org/user_builds/scmdata/checkouts/stable/src/scmdata/_xarray.py:236: FutureWarning: The previous implementation of stack is deprecated and will be removed in a future version of pandas. See the What's New notes for pandas 2.1.0 for details. Specify future_stack=True to adopt the new implementation and silence this warning.
else timeseries.T.stack(dimensions)
/home/docs/checkouts/readthedocs.org/user_builds/scmdata/checkouts/stable/src/scmdata/_xarray.py:236: FutureWarning: The previous implementation of stack is deprecated and will be removed in a future version of pandas. See the What's New notes for pandas 2.1.0 for details. Specify future_stack=True to adopt the new implementation and silence this warning.
else timeseries.T.stack(dimensions)
ScmRun.from_nc(OUTPUT_DIR / "out-ssp585-sparse.nc").filter(
variable="Surface Temperature"
).line_plot()
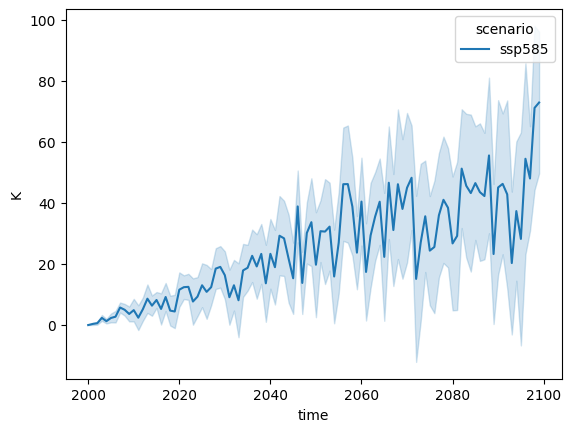
For such a data set, since both run_id and paraset_id vary, both could be added as dimensions
in the file.
The one problem with this approach is that you get very sparse arrays because the data is written on a 100 x 30 x 90 (time points x paraset_id x run_id) grid but there’s only 90 timeseries so you end up with 180 timeseries worth of nans (although this is a relatively small problem because the netCDF files use compression to minismise the impact of the extra nan values).
xr.load_dataset(OUTPUT_DIR / "out-ssp585-sparse.nc")
<xarray.Dataset>
Dimensions: (time: 100, run_id: 30, paraset_id: 10)
Coordinates:
* time (time) datetime64[ns] 2000-01-01 ... 209...
* run_id (run_id) int64 60 61 62 63 ... 86 87 88 89
* paraset_id (paraset_id) int64 0 1 2 3 4 5 6 7 8 9
Data variables:
Surface_Temperature (run_id, paraset_id, time) float64 0.0 ....
Atmospheric_Concentrations__CO2 (run_id, paraset_id, time) float64 nan ....
Radiative_Forcing (run_id, paraset_id, time) float64 nan ....
Attributes:
scmdata_metadata_scenario: ssp585
scmdata_metadata_model: example
scmdata_metadata_region: World
created_at: 2024-01-29T07:18:02.481132
_scmdata_version: 1.0.0# Load all scenarios
run_append([ScmRun.from_nc(fname) for fname in OUTPUT_DIR.glob("out-ssp*-sparse.nc")])
<ScmRun (timeseries: 90, timepoints: 100)>
Time:
Start: 2000-01-01T00:00:00
End: 2099-01-01T00:00:00
Meta:
model paraset_id region run_id scenario unit \
0 example 0 World 30 ssp370 K
1 example 0 World 31 ssp370 ppm
2 example 0 World 32 ssp370 W/m^2
3 example 1 World 33 ssp370 K
4 example 1 World 34 ssp370 ppm
.. ... ... ... ... ... ...
85 example 8 World 85 ssp585 ppm
86 example 8 World 86 ssp585 W/m^2
87 example 9 World 87 ssp585 K
88 example 9 World 88 ssp585 ppm
89 example 9 World 89 ssp585 W/m^2
variable
0 Surface Temperature
1 Atmospheric Concentrations|CO2
2 Radiative Forcing
3 Surface Temperature
4 Atmospheric Concentrations|CO2
.. ...
85 Atmospheric Concentrations|CO2
86 Radiative Forcing
87 Surface Temperature
88 Atmospheric Concentrations|CO2
89 Radiative Forcing
[90 rows x 7 columns]
An alternative to the sparse arrays is to specify the variables in the extras attribute. If
possible, this adds the metadata to the netCDF file as an extra co-ordinate, which uses one of
the dimensions as it’s co-ordinate. If using one of the dimensions as a co-ordinate would not
specify the metadata uniquely, we add the extra as an additional co-ordinate, which itself has
co-ordinates of _id. This _id co-ordinate provides a unique mapping between the extra metadata
and the timeseries.
for sce_run in large_run.groupby("scenario"):
sce = sce_run.get_unique_meta("scenario", True)
sce_run.to_nc(
OUTPUT_DIR / f"out-{sce}-extras.nc",
dimensions=["run_id"],
extras=["paraset_id"],
)
/home/docs/checkouts/readthedocs.org/user_builds/scmdata/checkouts/stable/src/scmdata/_xarray.py:201: FutureWarning: Series.__getitem__ treating keys as positions is deprecated. In a future version, integer keys will always be treated as labels (consistent with DataFrame behavior). To access a value by position, use `ser.iloc[pos]`
max_count = checker.groupby(col2).count().max()[0]
/home/docs/checkouts/readthedocs.org/user_builds/scmdata/checkouts/stable/src/scmdata/_xarray.py:236: FutureWarning: The previous implementation of stack is deprecated and will be removed in a future version of pandas. See the What's New notes for pandas 2.1.0 for details. Specify future_stack=True to adopt the new implementation and silence this warning.
else timeseries.T.stack(dimensions)
/home/docs/checkouts/readthedocs.org/user_builds/scmdata/checkouts/stable/src/scmdata/_xarray.py:201: FutureWarning: Series.__getitem__ treating keys as positions is deprecated. In a future version, integer keys will always be treated as labels (consistent with DataFrame behavior). To access a value by position, use `ser.iloc[pos]`
max_count = checker.groupby(col2).count().max()[0]
/home/docs/checkouts/readthedocs.org/user_builds/scmdata/checkouts/stable/src/scmdata/_xarray.py:236: FutureWarning: The previous implementation of stack is deprecated and will be removed in a future version of pandas. See the What's New notes for pandas 2.1.0 for details. Specify future_stack=True to adopt the new implementation and silence this warning.
else timeseries.T.stack(dimensions)
/home/docs/checkouts/readthedocs.org/user_builds/scmdata/checkouts/stable/src/scmdata/_xarray.py:201: FutureWarning: Series.__getitem__ treating keys as positions is deprecated. In a future version, integer keys will always be treated as labels (consistent with DataFrame behavior). To access a value by position, use `ser.iloc[pos]`
max_count = checker.groupby(col2).count().max()[0]
/home/docs/checkouts/readthedocs.org/user_builds/scmdata/checkouts/stable/src/scmdata/_xarray.py:236: FutureWarning: The previous implementation of stack is deprecated and will be removed in a future version of pandas. See the What's New notes for pandas 2.1.0 for details. Specify future_stack=True to adopt the new implementation and silence this warning.
else timeseries.T.stack(dimensions)
paraset_id is uniquely defined by run_id so we don’t end up with an extra _id co-ordinate.
xr.load_dataset(OUTPUT_DIR / "out-ssp585-extras.nc")
<xarray.Dataset>
Dimensions: (time: 100, run_id: 30)
Coordinates:
* time (time) datetime64[ns] 2000-01-01 ... 209...
* run_id (run_id) int64 60 61 62 63 ... 86 87 88 89
paraset_id (run_id) int64 0 0 0 1 1 1 ... 8 8 8 9 9 9
Data variables:
Surface_Temperature (run_id, time) float64 0.0 0.6824 ... nan
Atmospheric_Concentrations__CO2 (run_id, time) float64 nan nan ... nan nan
Radiative_Forcing (run_id, time) float64 nan nan ... 9.579
Attributes:
scmdata_metadata_scenario: ssp585
scmdata_metadata_model: example
scmdata_metadata_region: World
created_at: 2024-01-29T07:18:03.360352
_scmdata_version: 1.0.0ScmRun.from_nc(OUTPUT_DIR / "out-ssp585-extras.nc").filter(
variable="Surface Temperature"
).line_plot()

If we use dimensions and extra such that our extra co-ordinates are not uniquely defined by the
regions, an _id dimension is automatically added to ensure we don’t lose any information.
large_run.to_nc(
OUTPUT_DIR / "out-extras-sparse.nc",
dimensions=["scenario"],
extras=["paraset_id", "run_id"],
)
/home/docs/checkouts/readthedocs.org/user_builds/scmdata/checkouts/stable/src/scmdata/_xarray.py:201: FutureWarning: Series.__getitem__ treating keys as positions is deprecated. In a future version, integer keys will always be treated as labels (consistent with DataFrame behavior). To access a value by position, use `ser.iloc[pos]`
max_count = checker.groupby(col2).count().max()[0]
/home/docs/checkouts/readthedocs.org/user_builds/scmdata/checkouts/stable/src/scmdata/_xarray.py:201: FutureWarning: Series.__getitem__ treating keys as positions is deprecated. In a future version, integer keys will always be treated as labels (consistent with DataFrame behavior). To access a value by position, use `ser.iloc[pos]`
max_count = checker.groupby(col2).count().max()[0]
/home/docs/checkouts/readthedocs.org/user_builds/scmdata/checkouts/stable/src/scmdata/_xarray.py:234: FutureWarning: The previous implementation of stack is deprecated and will be removed in a future version of pandas. See the What's New notes for pandas 2.1.0 for details. Specify future_stack=True to adopt the new implementation and silence this warning.
timeseries.T.stack([*dimensions, "_id"])
xr.load_dataset(OUTPUT_DIR / "out-extras-sparse.nc")
<xarray.Dataset>
Dimensions: (time: 100, scenario: 3, _id: 90)
Coordinates:
* time (time) datetime64[ns] 2000-01-01 ... 209...
* scenario (scenario) <U6 'ssp119' 'ssp370' 'ssp585'
* _id (_id) int64 0 1 2 3 4 5 ... 85 86 87 88 89
paraset_id (_id) int64 0 0 0 1 1 1 2 ... 7 8 8 8 9 9 9
run_id (_id) int64 0 1 2 3 4 5 ... 85 86 87 88 89
Data variables:
Surface_Temperature (scenario, time, _id) float64 0.0 ... nan
Atmospheric_Concentrations__CO2 (scenario, time, _id) float64 nan ... nan
Radiative_Forcing (scenario, time, _id) float64 nan ... 9.579
Attributes:
scmdata_metadata_model: example
scmdata_metadata_region: World
created_at: 2024-01-29T07:18:04.000759
_scmdata_version: 1.0.0Multi-dimensional data
scmdata can also handle having more than one dimension. This can be especially helpful if you have output from a number of models (IAMs), scenarios, regions and runs.
multi_dimensional_run = []
for model in ["AIM", "GCAM", "MESSAGE", "REMIND"]:
for sce in ["ssp119", "ssp370", "ssp585"]:
for region in ["World", "R5LAM", "R5MAF", "R5ASIA", "R5OECD", "R5REF"]:
multi_dimensional_run.extend(
[
new_timeseries(
count=3,
model=model,
scenario=sce,
region=region,
variable=[
"Surface Temperature",
"Atmospheric Concentrations|CO2",
"Radiative Forcing",
],
unit=["K", "ppm", "W/m^2"],
paraset_id=paraset_id,
)
for paraset_id in range(10)
]
)
multi_dimensional_run = run_append(multi_dimensional_run)
multi_dimensional_run
<ScmRun (timeseries: 2160, timepoints: 100)>
Time:
Start: 2000-01-01T00:00:00
End: 2099-01-01T00:00:00
Meta:
model paraset_id region scenario unit \
0 AIM 0 World ssp119 K
1 AIM 0 World ssp119 ppm
2 AIM 0 World ssp119 W/m^2
3 AIM 1 World ssp119 K
4 AIM 1 World ssp119 ppm
... ... ... ... ... ...
2155 REMIND 8 R5REF ssp585 ppm
2156 REMIND 8 R5REF ssp585 W/m^2
2157 REMIND 9 R5REF ssp585 K
2158 REMIND 9 R5REF ssp585 ppm
2159 REMIND 9 R5REF ssp585 W/m^2
variable
0 Surface Temperature
1 Atmospheric Concentrations|CO2
2 Radiative Forcing
3 Surface Temperature
4 Atmospheric Concentrations|CO2
... ...
2155 Atmospheric Concentrations|CO2
2156 Radiative Forcing
2157 Surface Temperature
2158 Atmospheric Concentrations|CO2
2159 Radiative Forcing
[2160 rows x 6 columns]
multi_dim_outfile = OUTPUT_DIR / "out-multi-dimensional.nc"
multi_dimensional_run.to_nc(
multi_dim_outfile,
dimensions=("region", "model", "scenario", "paraset_id"),
)
/home/docs/checkouts/readthedocs.org/user_builds/scmdata/checkouts/stable/src/scmdata/_xarray.py:236: FutureWarning: The previous implementation of stack is deprecated and will be removed in a future version of pandas. See the What's New notes for pandas 2.1.0 for details. Specify future_stack=True to adopt the new implementation and silence this warning.
else timeseries.T.stack(dimensions)
xr.load_dataset(multi_dim_outfile)
<xarray.Dataset>
Dimensions: (time: 100, scenario: 3, region: 6,
paraset_id: 10, model: 4)
Coordinates:
* time (time) datetime64[ns] 2000-01-01 ... 209...
* scenario (scenario) <U6 'ssp119' 'ssp370' 'ssp585'
* region (region) <U6 'R5ASIA' 'R5LAM' ... 'World'
* paraset_id (paraset_id) int64 0 1 2 3 4 5 6 7 8 9
* model (model) <U7 'AIM' 'GCAM' 'MESSAGE' 'REMIND'
Data variables:
Surface_Temperature (region, model, scenario, paraset_id, time) float64 ...
Atmospheric_Concentrations__CO2 (region, model, scenario, paraset_id, time) float64 ...
Radiative_Forcing (region, model, scenario, paraset_id, time) float64 ...
Attributes:
created_at: 2024-01-29T07:18:09.510062
_scmdata_version: 1.0.0multi_dim_loaded_co2_conc = ScmRun.from_nc(multi_dim_outfile).filter(
variable="Atmospheric Concentrations|CO2"
)
seaborn_df = multi_dim_loaded_co2_conc.long_data()
seaborn_df.head()
| model | paraset_id | region | scenario | unit | variable | time | value | |
|---|---|---|---|---|---|---|---|---|
| 0 | AIM | 0 | R5ASIA | ssp119 | ppm | Atmospheric Concentrations|CO2 | 2000-01-01 | 0.000000 |
| 1 | AIM | 0 | R5ASIA | ssp119 | ppm | Atmospheric Concentrations|CO2 | 2001-01-01 | 0.730551 |
| 2 | AIM | 0 | R5ASIA | ssp119 | ppm | Atmospheric Concentrations|CO2 | 2002-01-01 | 0.142061 |
| 3 | AIM | 0 | R5ASIA | ssp119 | ppm | Atmospheric Concentrations|CO2 | 2003-01-01 | 2.977609 |
| 4 | AIM | 0 | R5ASIA | ssp119 | ppm | Atmospheric Concentrations|CO2 | 2004-01-01 | 0.785949 |
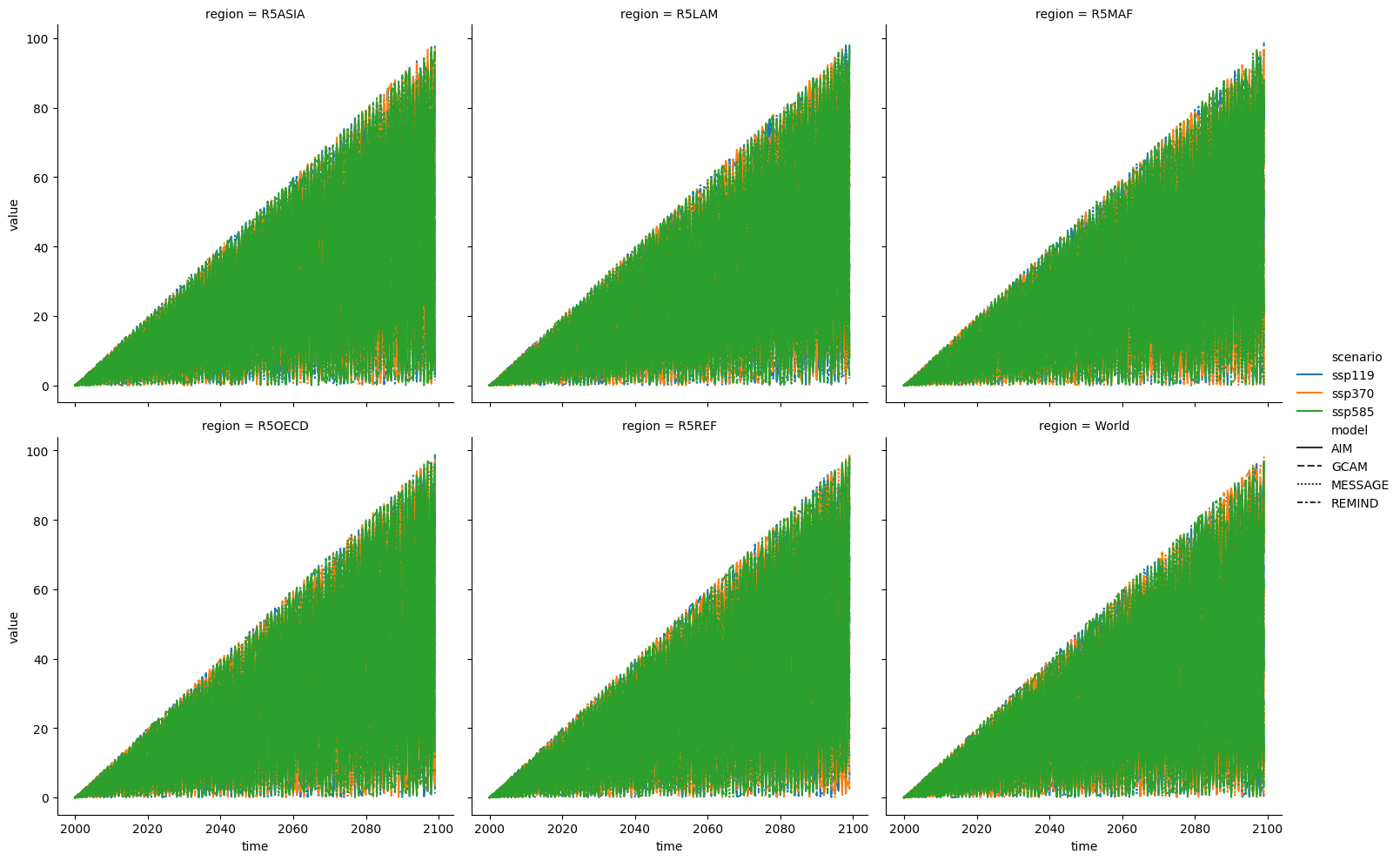
sns.relplot(
data=seaborn_df,
x="time",
y="value",
units="paraset_id",
estimator=None,
hue="scenario",
style="model",
col="region",
col_wrap=3,
kind="line",
)
<seaborn.axisgrid.FacetGrid at 0x7f353ebbc490>